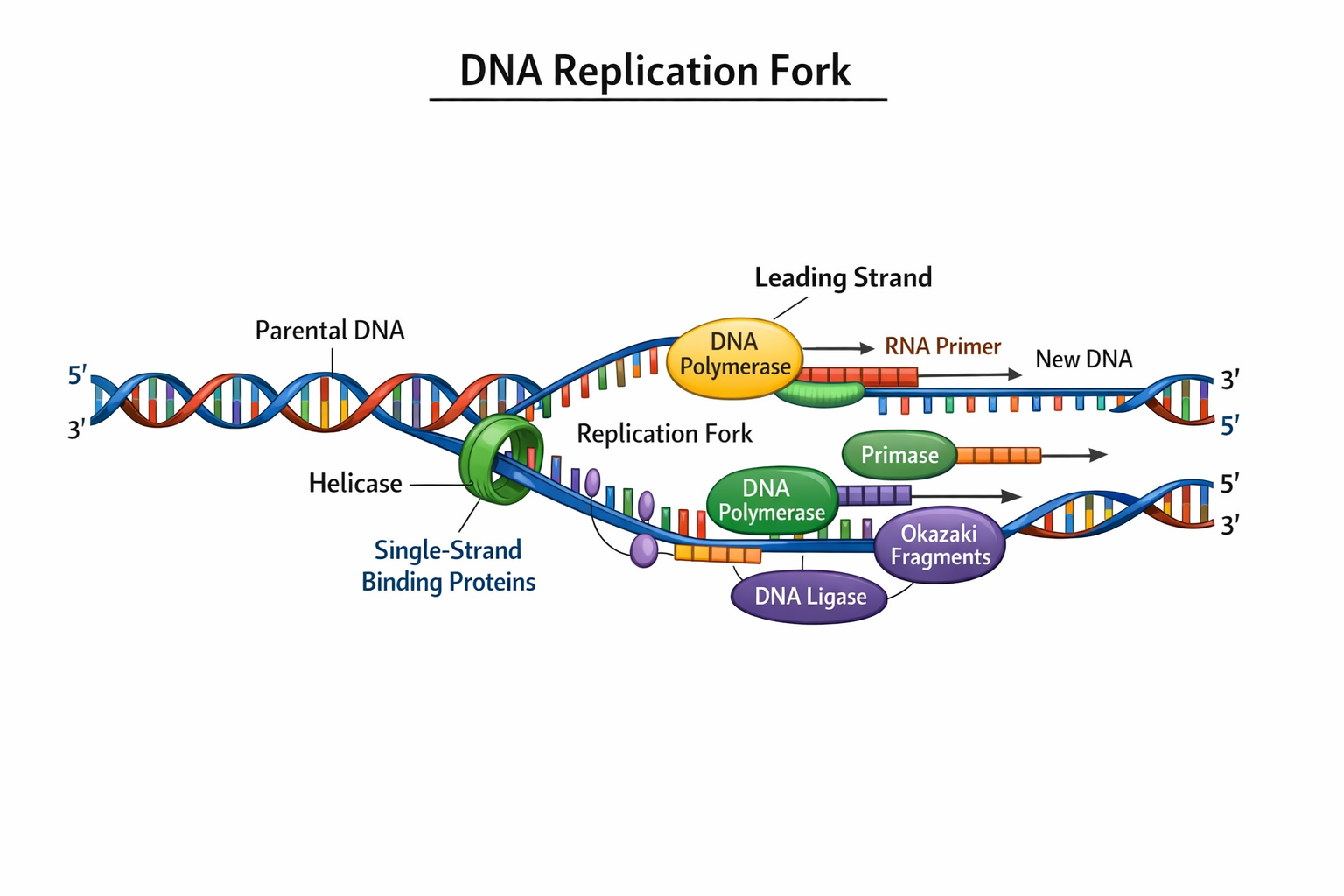
DNA Replication Fork
DNA Replication Fork : Labeled Diagram, Function, and Definition
Explore what is a replication fork, see a labeled DNA replication fork diagram, and understand where the replication fork is in the genome. Learn its key role in DNA synthesis.
What Is a Replication Fork?
The replication fork is the Y-shaped region of DNA where the double helix unwinds, allowing DNA polymerase to synthesize new complementary strands. This site marks the active area where DNA replication is happening.
During DNA replication, the double helix unwinds and replication proceeds in both directions from the origin of replication, creating two replication forks that move away from each other.
Replication Fork Diagram
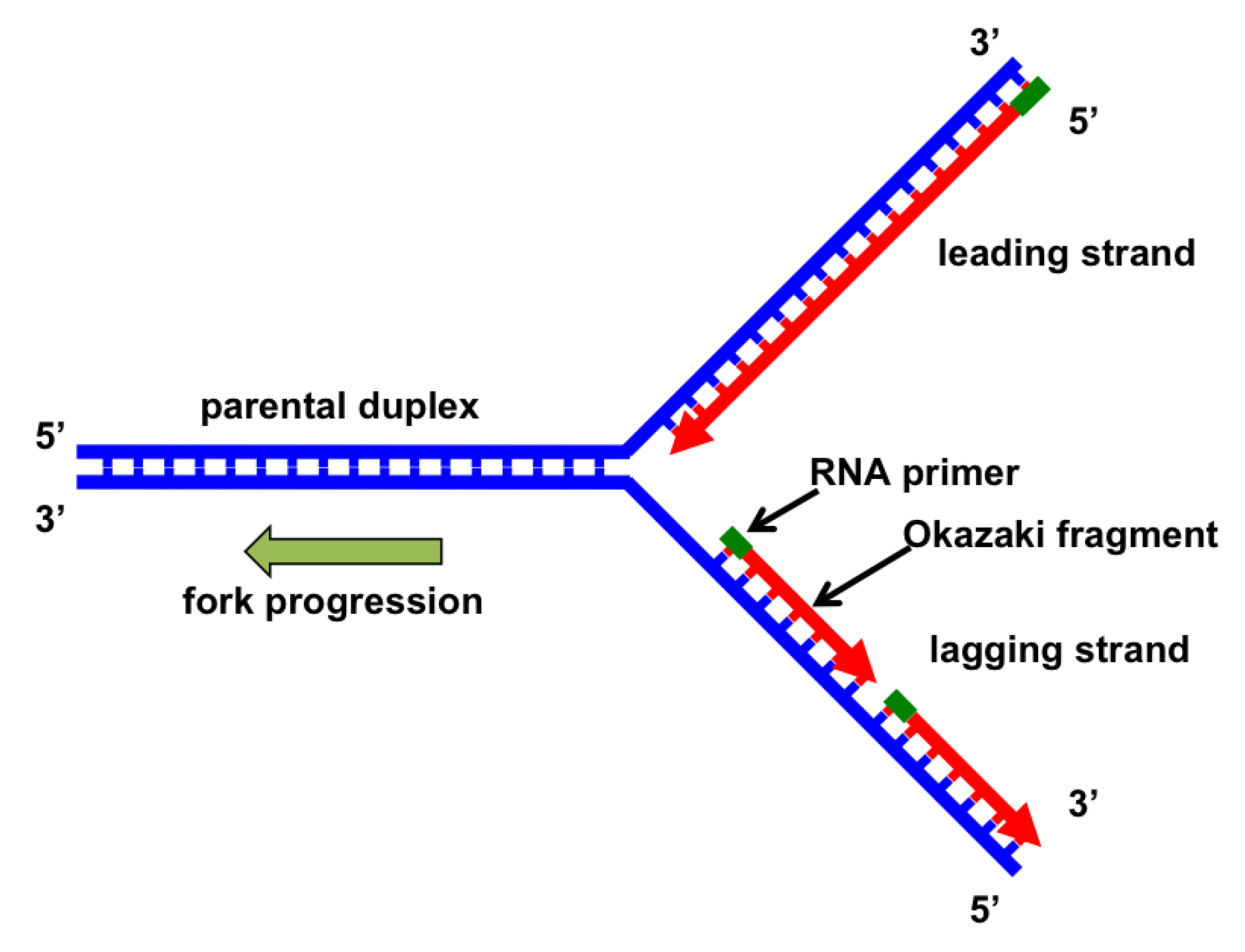
DNA Replication Fork Structure and Function
At each DNA replication fork, two strands are produced:
- The leading strand is synthesized continuously.
- The lagging strand is synthesized in short fragments called Okazaki fragments.
Multiple enzymes operate here:
- Helicase unwinds the DNA.
- Primase lays RNA primers.
- DNA polymerase extends the new strands.
- Ligase joins lagging strand fragments.
These enzymes work in a coordinated complex that drives semiconservative replication.
Labeled DNA Replication Fork (Description)Below is a clear description of the main parts of a replication fork that you can use to label your diagram:
1- Parental DNA : the original DNA double helix before unwinding.
2- Replication Fork : the active area where the DNA is split into two single strands.
3- Leading Strand : the new complementary strand synthesized continuously in the 5′ → 3′ direction.
4- Lagging Strand : the new complementary strand synthesized discontinuously in short segments.
5- Okazaki Fragments : short DNA pieces on the lagging strand which are later joined together.
6- DNA Helicase : the enzyme that unwinds and separates the two DNA strands.
7- Single‑Strand Binding Proteins (SSBs) : proteins that stabilize single strands and prevent them from snapping back together.
8- DNA Polymerase : the enzyme that synthesizes new DNA strands by adding nucleotides.
9- RNA Primer : short RNA sequence that provides a starting point for DNA synthesis.
10- DNA Ligase : the enzyme that joins Okazaki fragments into a continuous strand.
Replication Fork in Action
At each replication fork :
- Helicase unwinds the DNA double helix.
- The two separated strands act as templates.
- On the leading strand, DNA polymerase adds nucleotides continuously.
- On the lagging strand, DNA is synthesized in short sections called Okazaki fragments.
- RNA primers are later removed and replaced with DNA.
- DNA ligase then links the Okazaki fragments to form a complete strand.
Key Features & Functions
| Feature | Role |
|---|---|
| Replication Fork => | Active region of DNA unwinding and synthesis |
| Helicase => | Opens up the DNA helix |
| DNA Polymerase => | Builds new DNA strands |
| SSB Proteins => | Stabilize separated DNA strands |
| Okazaki Fragments => | Pieces of new DNA on lagging strand |
| DNA Ligase => | Seals gaps between fragments |
Where Is the Replication Fork Located?
The replication fork forms at origins of replication, which are specific sequences where the unwinding of DNA begins. In:
- Eukaryotic cells : multiple replication forks form simultaneously along linear chromosomes. Pubmed
- Prokaryotic cells : typically one origin forms two forks that progress in opposite directions.

DNA Replication Fork Animation
During DNA replication inside a cell, each of the two old DNA strands serves as a template for the formation of an entire new strand. Because each of the to daughters of a dividing cell inherits a new DNA double helix containing one old and one new strand, the DNA double helix is said to be replicated “semiconservetively” by DNA polymerase.
Analyses carried out in the early 1960s on whole replicating chromosomes revealed a localized region of replication that moves progressively along the parental DNA double helix. Because of its Y shaped structure, this reactive region is called a replication fork. At a replication fork, the DNA both new daughter strand is synthesized by a multienzyme complex that contains the DNA polymerase.
Summary Table: DNA Replication Fork Explained
| Term | Description |
|---|---|
| Replication fork | Y-shaped structure formed during DNA unwinding |
| DNA replication fork | Specific region where DNA is duplicated by polymerase |
| Replication fork diagram | Visual representation of replication process and enzyme positions |
| Replication fork labeled | Annotated diagram showing helicase, primase, polymerase, ligase, etc. |
| Where is the replication fork | Found at every origin of DNA synthesis during S-phase |
| Replication forks | Multiple forks occur simultaneously in eukaryotic chromosomes |
DNA Replication Fork Diagram Labeled
Common FAQ
Q: Why are two replication forks formed?
A: In circular and linear DNA, replication begins at an origin and proceeds bidirectionally, forming two forks that increase replication speed.
Q: What direction do DNA polymerases work in?
A: DNA polymerases add new bases only in the 5′ → 3′ direction.
Q: Why is the lagging strand synthesized in fragments?
A: Because DNA polymerase can’t follow the unwinding fork continuously in the 3′ → 5′ direction, short fragments are synthesized instead.
There are no products listed under this category.